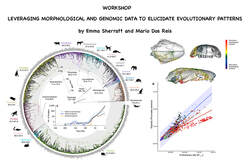
The workshop will be on Monday 1 May – Tuesday 2 May 2023 in Canberra ACT Australia.
Details:
In this workshop, participants will gain a deeper understanding of the theoretical background and practical applications of both morphological and genomic data in modern evolutionary studies.
The workshop covers the collection of morphometric data using microCT scans and explores its relevance in macroevolutionary studies. Additionally, participants will learn how to integrate this data into molecular analysis to infer speciation times, morphological rates of evolution, and ancestral character reconstructions.
Led by two leading experts in the field, Emma Sherratt (The University of Adelaide) and Mario Dos Reis (Queen Mary University of London, CSIRO visitor), this workshop includes two seminars and two practical sessions.
This is an opportunity to enhance your knowledge and skills in generating and integrating morphological and genomic data in phylogenetic studies.
 RSS Feed
RSS Feed
